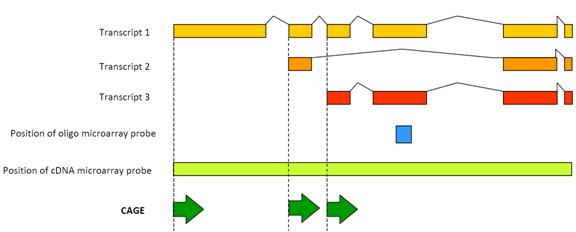
The miR-430 locus with extreme promoter density forms a transcription body during the minor wave of zygotic genome activation - ScienceDirect
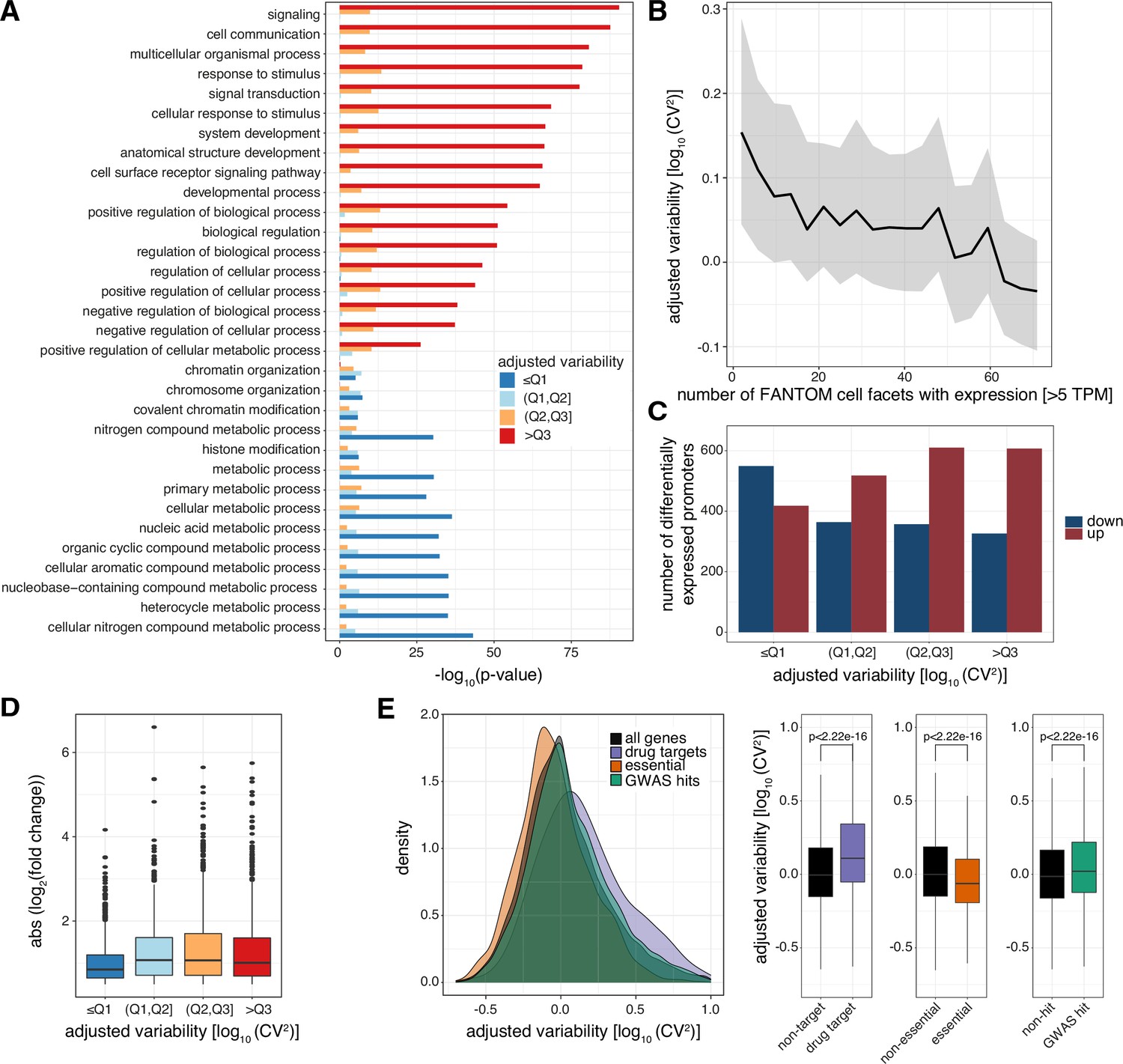
Promoter sequence and architecture determine expression variability and confer robustness to genetic variants | eLife
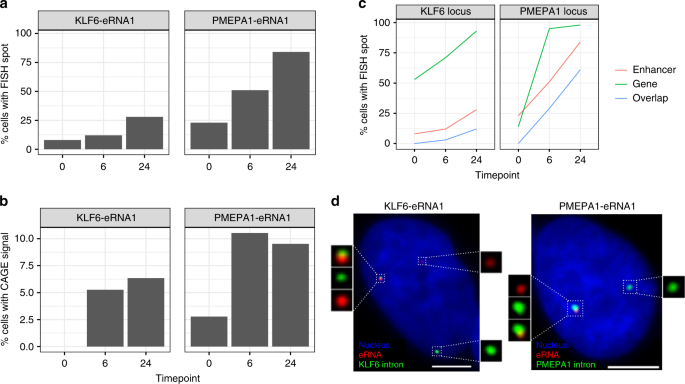
C1 CAGE detects transcription start sites and enhancer activity at single-cell resolution | Nature Communications
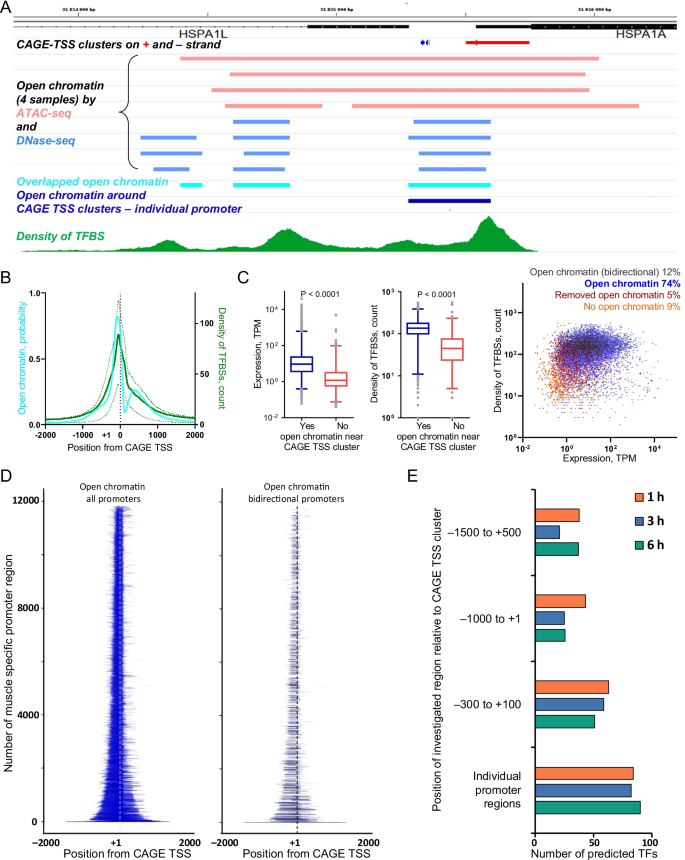
Alternative transcription start sites contribute to acute-stress-induced transcriptome response in human skeletal muscle | Human Genomics | Full Text
Influence of Rotational Nucleosome Positioning on Transcription Start Site Selection in Animal Promoters | PLOS Computational Biology

Multiple transcription initiation and alternative promoter usage in G.... | Download Scientific Diagram

Cap analysis of gene expression (CAGE) sequencing reveals alternative promoter usage in complex disease | bioRxiv

Cap analysis of gene expression reveals alternative promoter usage in a rat model of hypertension | Life Science Alliance
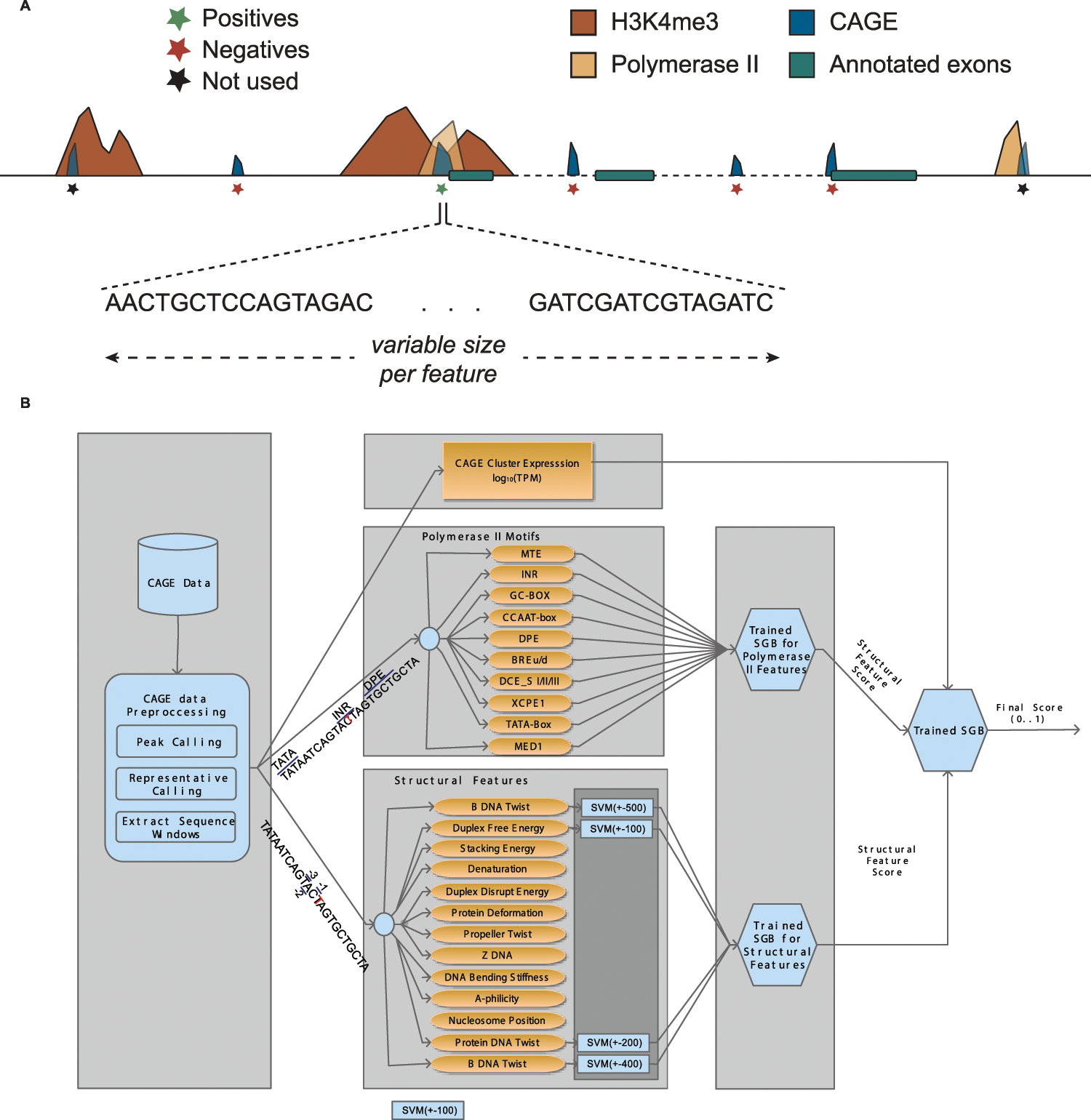
Solving the transcription start site identification problem with ADAPT-CAGE: a Machine Learning algorithm for the analysis of CAGE data | Scientific Reports

Promoter sequence and architecture determine expression variability and confer robustness to genetic variants | bioRxiv

Alternative promoters in CpG depleted regions pervasively account for epigenetic misregulation of cancer transcriptomes | bioRxiv
Distal CpG islands can serve as alternative promoters to transcribe genes with silenced proximal promoters
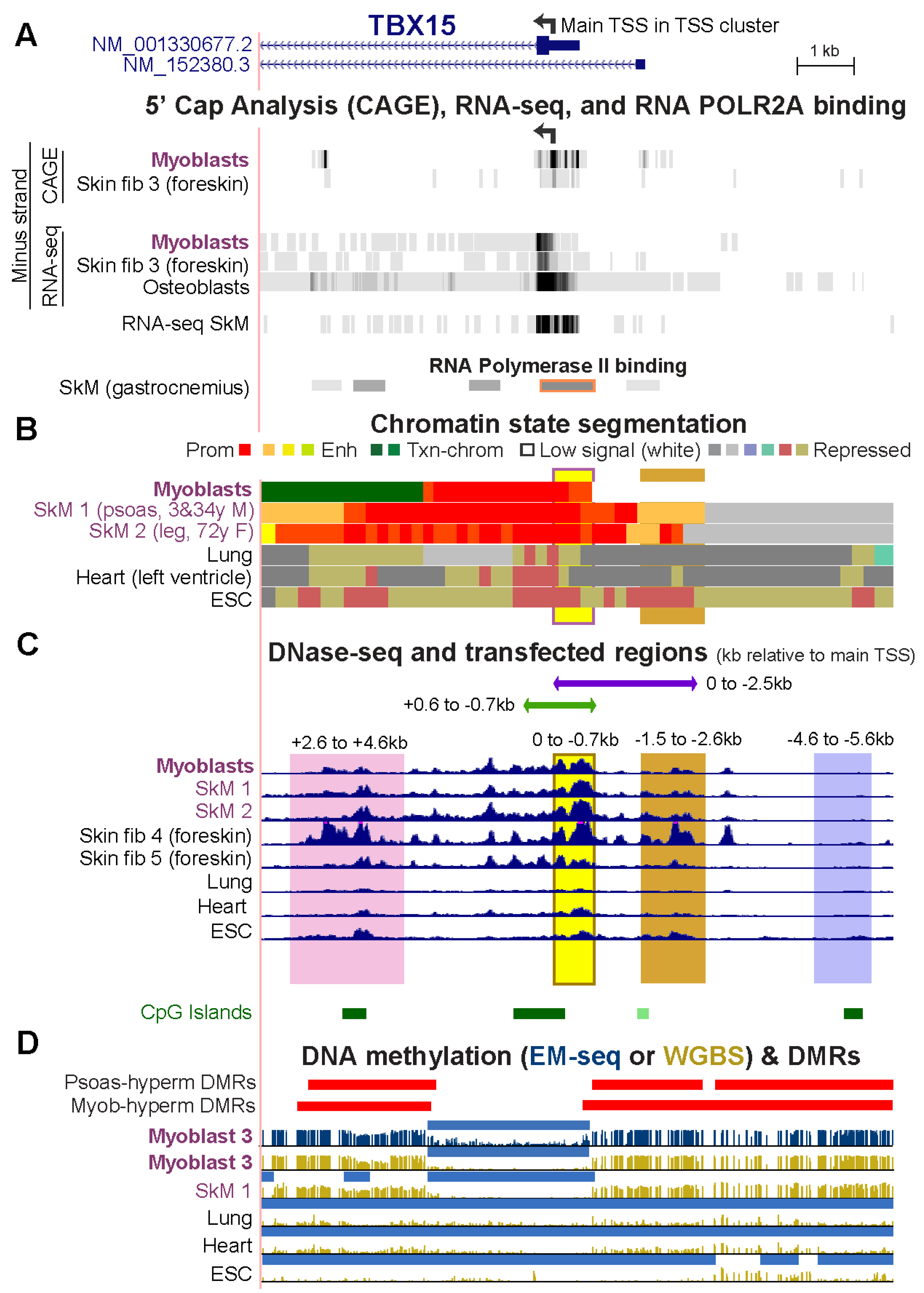
Epigenomes | Free Full-Text | Promoter-Adjacent DNA Hypermethylation Can Downmodulate Gene Expression: TBX15 in the Muscle Lineage